We provide Alphafold on-demand service running on Kubernetes. Alphafold is available at https://alphafold.cloud.e-infra.cz, sign with e-INFRA CZ AAI or LifeScience AAI.
Application is based on Jupyer notebook, therefore notebook loader is shown after login. It may take several minutes before Alphafold is ready to run.
Alphafold Parameters
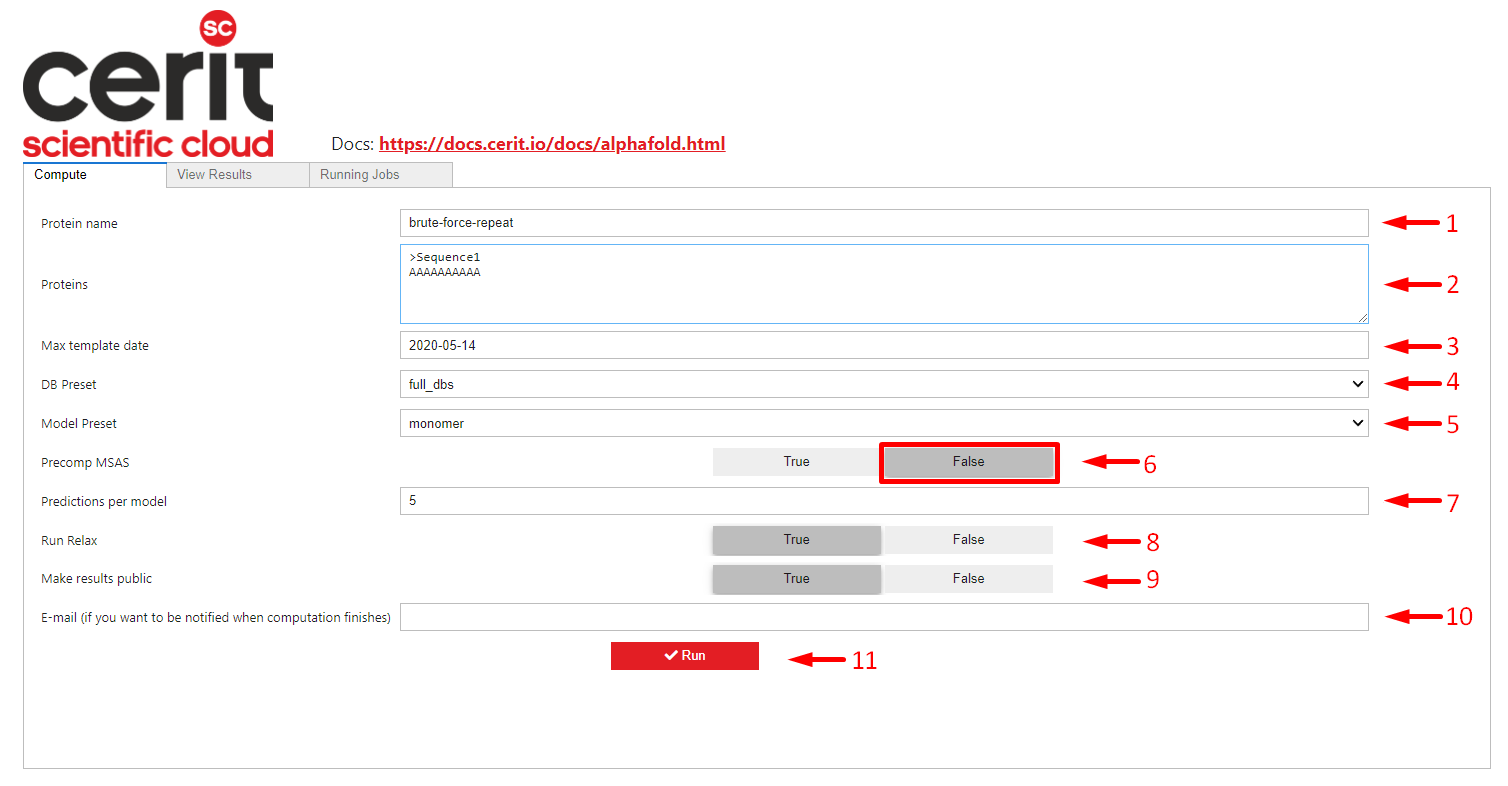
After successful login you will see screen as shown below. You need to fill parameters for Alphafold calculation, at least Proteins (arrow 2) is required, everything else can be left default. See here for additional parameters. Protein name (arrow 1) can be changed, but must be unique (you cannot run multiple computations with the same name), otherwise you will see the message: A job with this name already exists, please enter a different name.
By default, all calculation results are public, i.e. everyone who is able to log in can see them, but the name of the user who initiated the calculation is not shown. If this is not desired, publication can be disabled (arrow 9).
If you fill in email (arrow 10), you will receive a notification when the calculation is finished.
Run (arrow 11) starts the calculation.
Alphafold Running
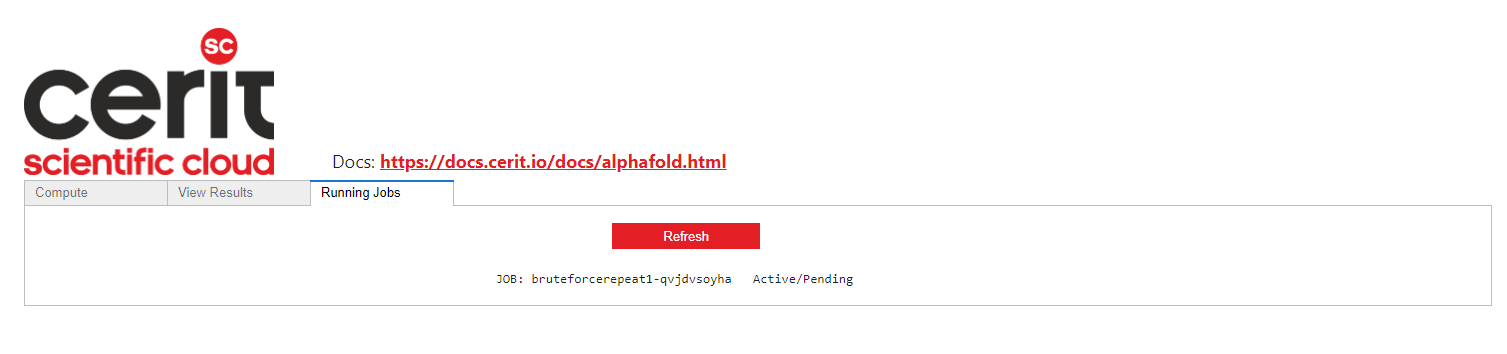
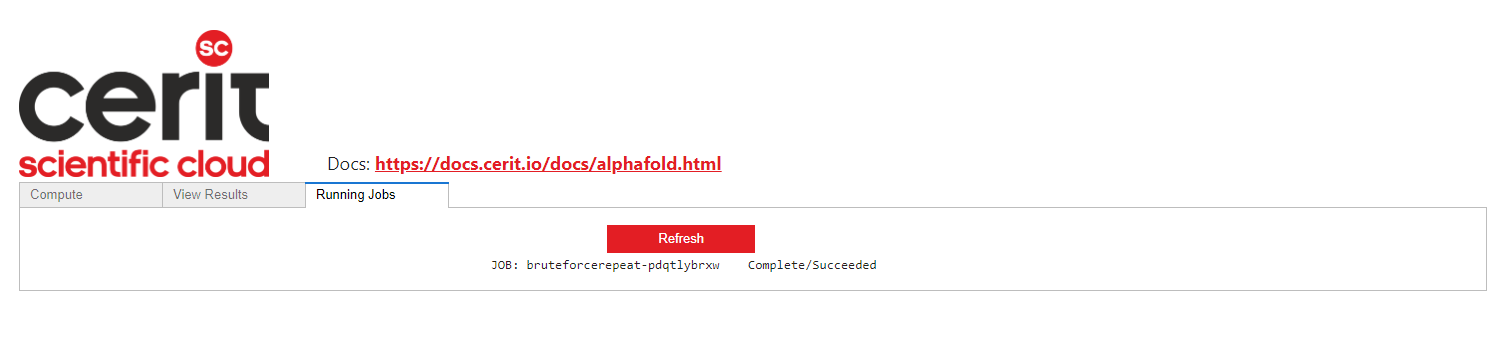
If a job has been submitted, it can be seen in the Running Jobs tab. It can be Pending, Running or Succeeded.
Viewing Results
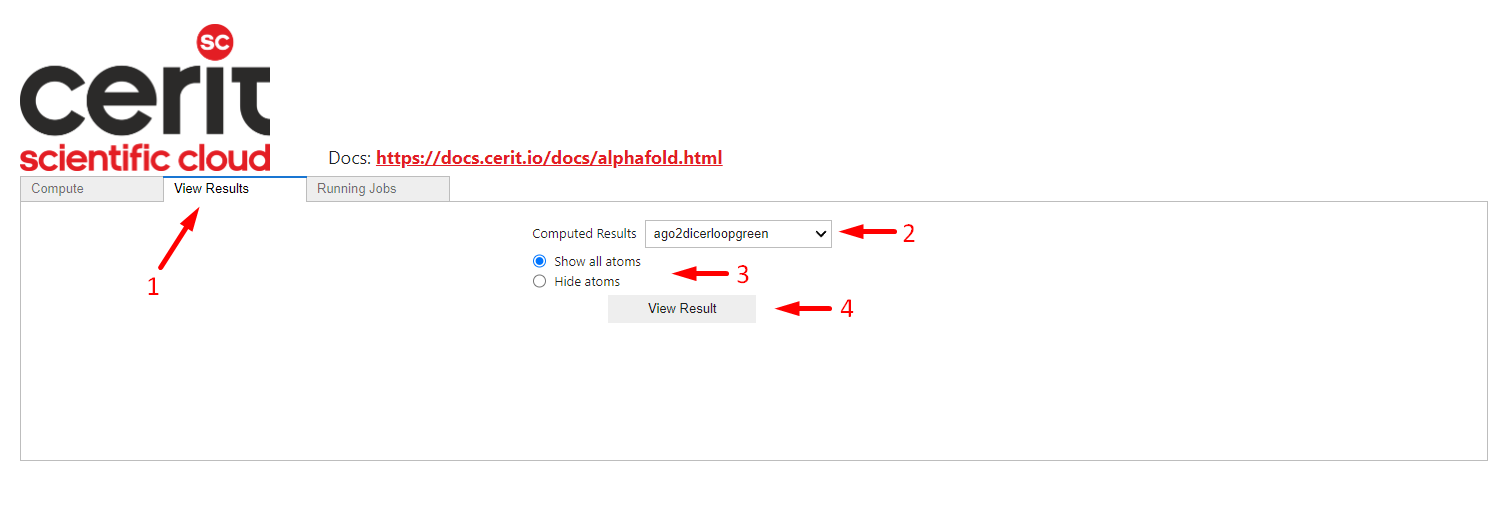
This application allows you to preview the calculated results in the View Results tab. You can find your protein name in selector (arrow 2), you can choose basic display parameters – whether to show atoms or not (arrow 3). The View Result button (4) displays the result in the browser. Viewer is interactive and allows rotation and panning.
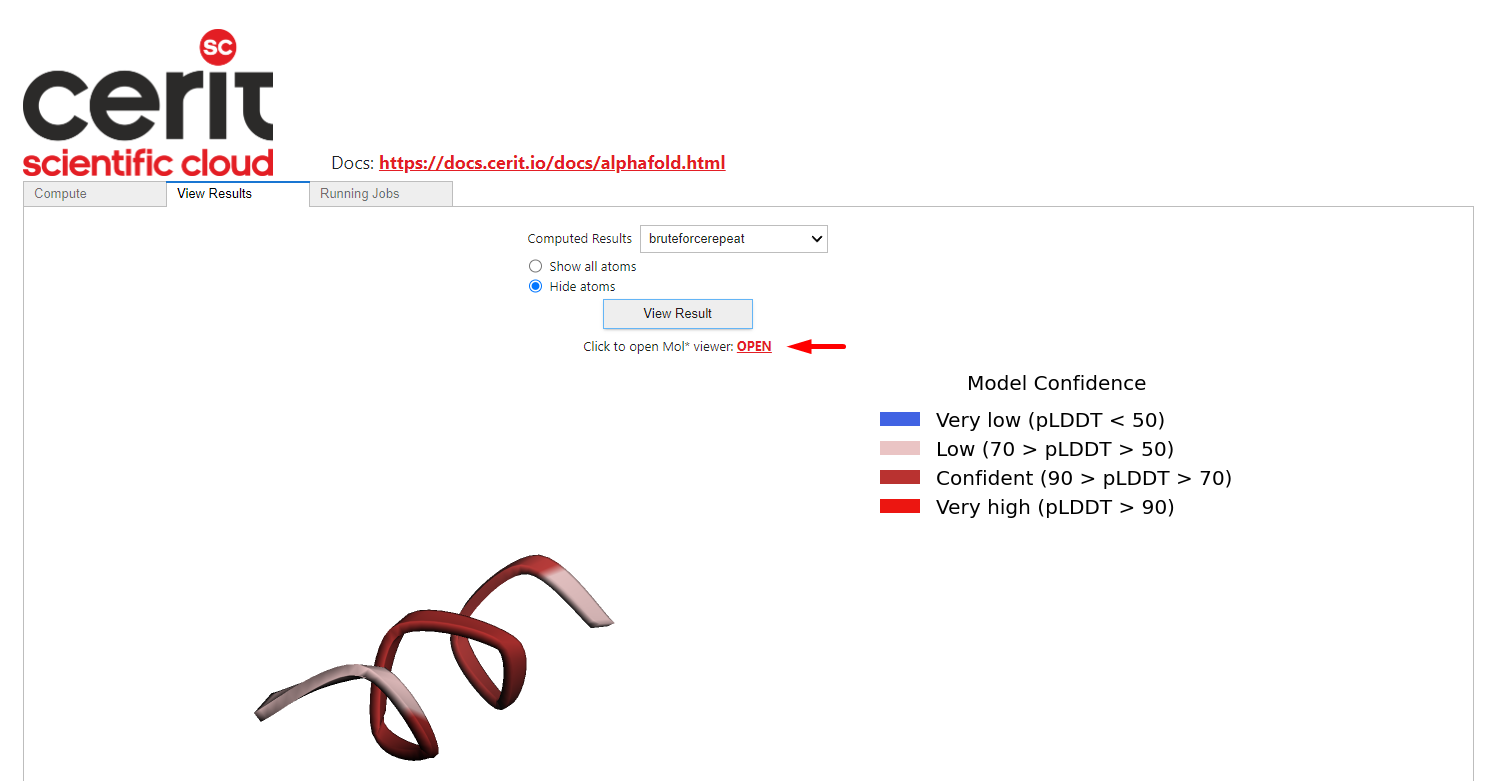
We also provide a more complex viewer – Mol* clicking on the OPEN link.
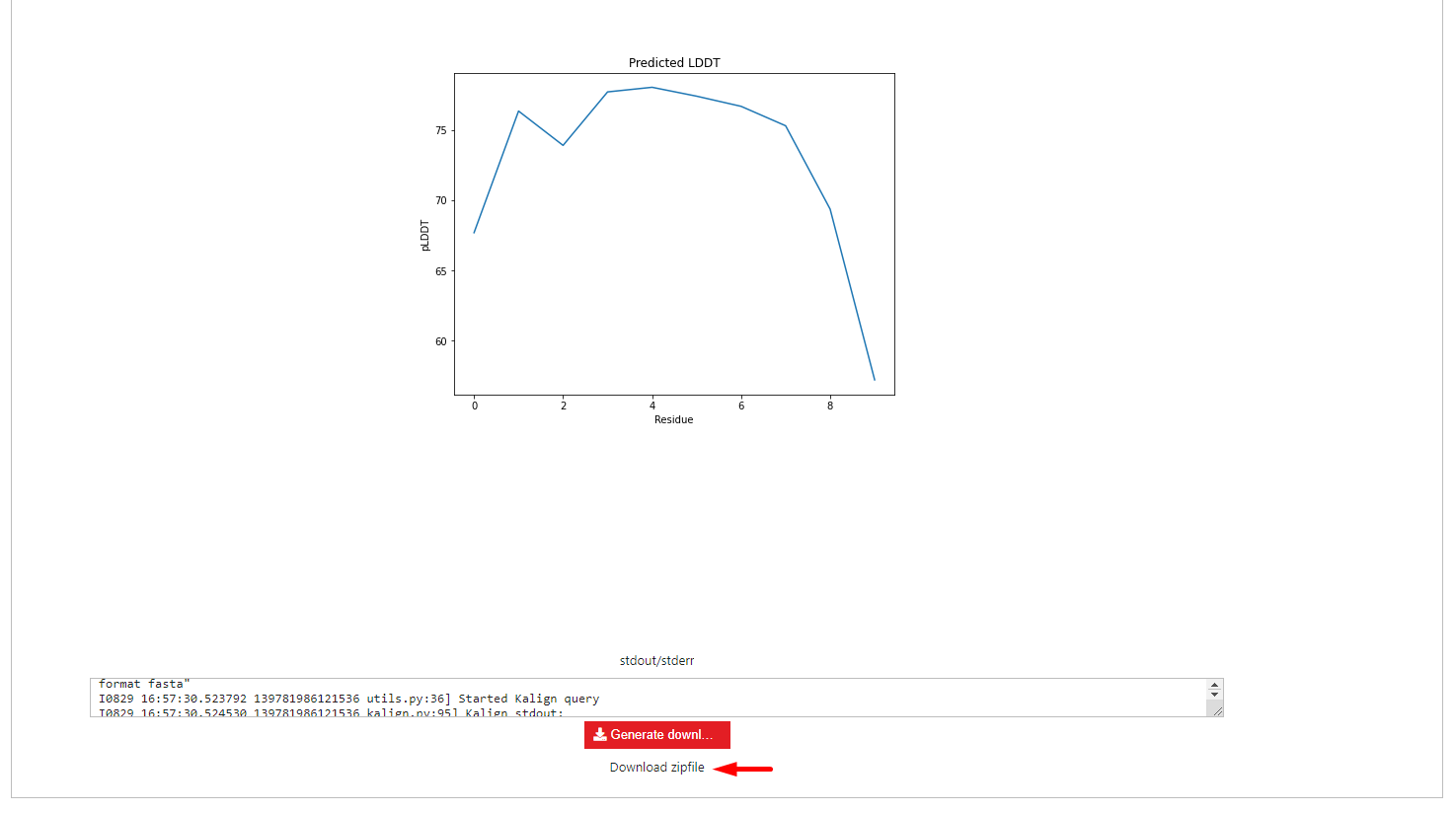
The results can be downloaded at the bottom of this page by clicking the Generate download button and following the download zipfile link.